Mauri Kostiainen has received a two million euro grant to study new biohybrid materials
The work of Mauri Kostiainen can help combine the best characteristics of biomolecules and synthetic materials.
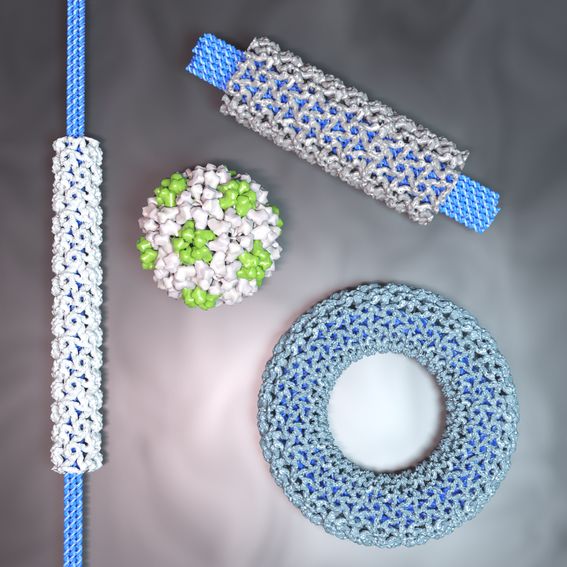

Bioengineers have found a way to program the size and shape of virus particles by combining viral protein building blocks and templates made from DNA. The resulting nanostructures could have applications in vaccine development and transporting drugs inside the body.
Virus capsid proteins—the proteins that shield the genome of a virus—can be used to build precisely structured protein assemblies. Their shapes and geometry, however, depend largely on the virus strain. Reprogramming these assemblies, no matter the original viral blueprint, is an intriguing possibility for drug delivery and vaccine development.
Scientists tackled the challenge by generating a “structured genome” template on which capsid proteins can assemble. To avoid deforming the flexible genome and creating unintended shapes, they used rigid DNA origami structures. These structures are only tens to hundreds of nanometres in length, but entirely made of DNA, which is folded accurately into the desired template shape.
‘Our approach is based on electrostatic interactions between the negative charge of the DNA nanostructures and a positively charged domain of the capsid proteins, paired with intrinsic interactions between the single proteins. By altering the amount of protein used, we can fine-tune the number of highly-ordered protein layers, which encapsulate the DNA origami,’ says Iris Seitz, lead author and doctoral researcher at Aalto University.
‘By using DNA origami as a template, we can direct the capsid proteins into a user-defined size and shape, resulting in assemblies which are well-defined, both in length and diameter. By testing a variety of DNA origami structures, we also learned how the templates’ geometry affected the whole assembly,’ Seitz adds.
‘With the help of cryogenic electron microscopy imaging, we were able to visualise the highly ordered proteins upon assembly and, with that, measure even small changes in the geometry of the assembly arising from different templates,’ explains professor Juha Huiskonen, a collaborating scientist from the University of Helsinki.
‘We have found a simple but effective strategy to (re)direct capsid proteins to a desired shape. Our approach is adaptable and therefore not limited to a single capsid protein type, as we demonstrated with capsid proteins from four different viruses. Additionally, we can tweak our template to be more application-relevant, for instance by integrating RNA into the origami, which could subsequently be translated into useful or site-specific proteins,’ explains Aalto professor Mauri Kostiainen, leader of the research project.
Although DNA origami structures are a promising material for interfacing biological systems, they suffer from instability, especially in the presence of DNA-degrading enzymes.
In experiments, however, ‘we can clearly observe that the protein layer efficiently protects the encapsulated DNA nanostructures from degradation. By combining protection with the functional properties of nucleic acid origami, including the possibility to deliver DNA or messenger RNA together with other cargo molecules, we believe that our approach provides interesting future directions for biomedical engineering,’ concludes Kostiainen.
This work was conducted jointly at Aalto University (Finland) with researchers from the University of Helsinki (Finland), Griffith University (Australia), Tampere University (Finland) and University of Twente (The Netherlands).
Reference:
Seitz et al. (2023). DNA-origami-directed virus capsid polymorphism. Nature Nanotechnology, doi: 10.1038/s41565-023-01443-x
Iris Seitz, doctoral researcher
iris.seitz@aalto.fi
Mauri Kostiainen, professor
mauri.kostiainen@aalto.fi
phone +358503627070

The work of Mauri Kostiainen can help combine the best characteristics of biomolecules and synthetic materials.
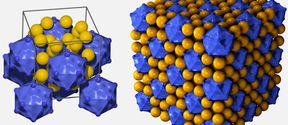
Group led by Professor Mauri Kostiainen

Gold nanoparticles are arranged by custom DNA molecules to produce colours

Electronically controlled molecular machines would be faster as well as easier to manufacture, as they would not need to rely on sophisticated chemical synthesis.

Nanosized hinges can fold and unfold on command

Understanding of computer science and mathematics becomes increasingly important in the field of DNA nanotechnology, says Professor Pekka Orponen

To DNA nanotechnologists, DNA is a smart building material that can be useful in the development of medical applications

DNA microscopy makes it possible to view biological molecules on micro-level without expensive optics.

Three-dimensional nano-sized structures obtained from DNA using a new design method


