Bioeconomy Infrastructure (external link)
Joint research infrastructure of Aalto University and VTT Bioeconomy infrastructure on Finland’s roadmap for research infrastructures 2021–2024
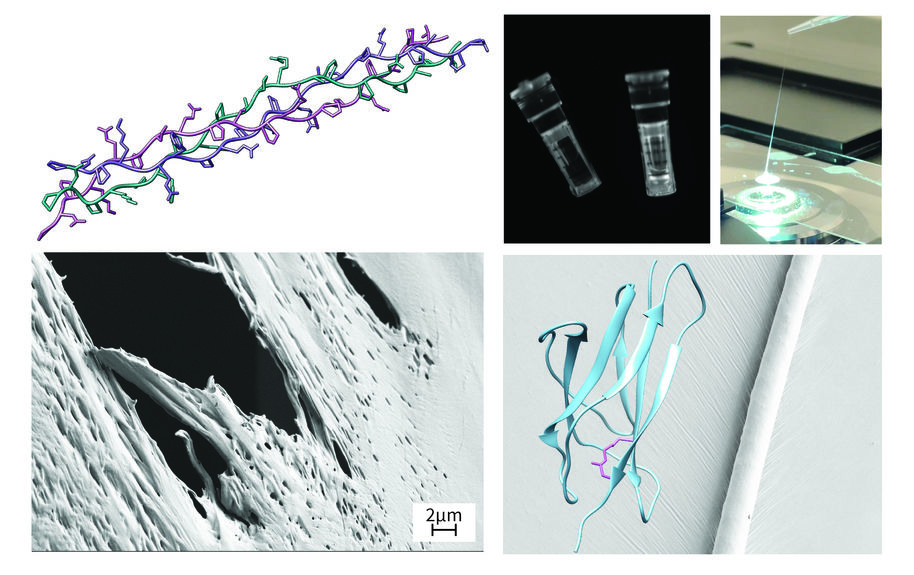
In this project, funded by the Research Council of Finland (formerly the Academy of Finland), we aim to improve the water resistance of currently available silk- and cellulose-based biomaterials by mimicking the crosslinking of natural silks and leather. The goal is to design fully biodegradable materials with outstanding mechanical properties in both dry and wet states. We hope that the novel crosslinked silk-CNF composite materials developed in the project can be used as a renewable and biodegradable alternative to replace current plastic- and animal-based materials.
https://research.aalto.fi/en/projects/crosssilk
The goal of this project, funded by the Novo Nordisk Foundation Emerging Investigator grant, is to develop methods to produce post-translationally modified structural proteins, including silks and collagens, in bacteria that cannot make the desired modifications naturally. There is accumulating evidence that the post-translational modifications of proteins are essential for the mechanical properties and functionalities of the resulting materials. Yet, the mechanisms behind how these modifications affect the properties of biomaterials are not fully understood, mainly due to technical limitations. Production in bacteria enables obtaining the proteins in an economical, ethical, and sustainable manner. We use the modified proteins to engineer novel functional biomaterials, which have the potential to substitute current oil- and animal-based alternatives.






Contact: sesilja.aranko@aalto.fi

Joint research infrastructure of Aalto University and VTT Bioeconomy infrastructure on Finland’s roadmap for research infrastructures 2021–2024

OtaNano is Finland's national research infrastructure for micro-, nano-, and quantum technologies

National fundamental research centre in Hamburg, Germany

With 28 member states, EMBL has more than 110 independent research groups and service teams covering the spectrum of molecular biology at six sites in Barcelona, Grenoble, Hamburg, Heidelberg, EMBL-EBI Hinxton, and Rome.

Staff Scientist Sesilja Aranko has been awarded the highly competed Emerging Investigator 2023 Grant from Novo Nordisk Foundation for a research project focusing on post-translational modifications in protein-based biomaterials. The grant awarded is 10 million DKK (approx. 1.34M€) for a five-year period.