PHYS brown bag seminar: Prokop Hapala (SIN): Computer aided AFM imaging & recognition of 3D molecules
Join us for science and pizza in the Nanotalo lobby.

When
Where
Event language(s)
Join us for science and pizza in the Nanotalo lobby!
Presentation by Prokop Hapala (SIN group, Department of Applied Physics, Aalto University)
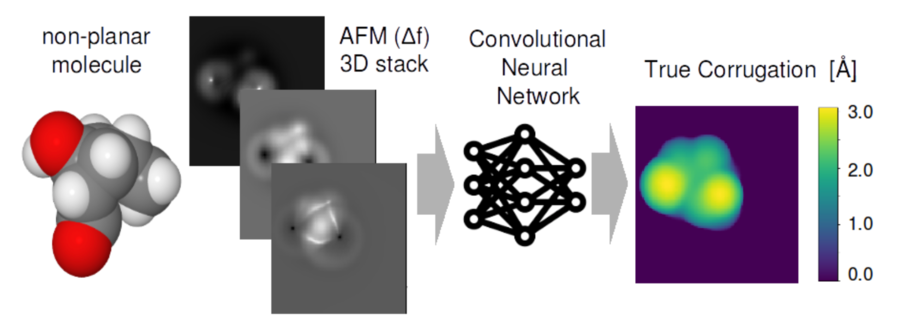
Abstract: In recent decades Atomic Force Microscopy with the tip functionalized by carbon monoxide (CO) has provided a unique tool to experimentally image sub-molecular details of individual organic molecules [1], which is of great importance e.g. for on-surface chemistry. Most experiments have, however, up to now been limited to flat aromatic molecules, due to difficulties in interpreting highly distorted AFM images originating from non-planer molecules and due to mechanical relaxation of the tip or sample. These problems can be partially overcome using a simple mechanical model (Probe-Particle Model [2]) which can reproduce those distortions and therefore simulate AFM images for a given molecular structure. However, this still requires a laborious search for the molecular structure that reproduces that particular experimental image. We instead attempt to develop automatic tools to conduct the inverse task – to recover molecular structure from a given set of AFM images. Preliminary results suggest that convolutional neural network (CNN) [3] trained on simulated AFM images can learn this inverse mapping rather easily. Yet application of the method on real experimental data, and identification of atomic species remains a challenge.
[1] Gross, L., Mohn, F., Moll, N., Liljeroth, P., & Meyer, G.; The chemical structure of a molecule resolved by atomic force microscopy. Science (New York, N.Y.), 325(5944), 1110–1114 (2009).
[2] Hapala, P., Kichin, G., Wagner, C., Tautz, F. S., Temirov, R., & Jelínek, P. Physical Review B, 90(8), 085421 (2014).
[3] Lecun, Y., Bottou, L., Bengio, Y., & Haffner, P.; Proceedings of the IEEE, 86(11), 2278–2324 (1998).






